Difference between revisions of "Dx map"
m (Switch to the {{gna bug url}} and {{gna bug url}} templates to remove dead Gna! links.) |
|||
| (8 intermediate revisions by 3 users not shown) | |||
| Line 1: | Line 1: | ||
| + | __TOC__ | ||
| + | |||
| + | == Sample data to try dx == | ||
| + | <source lang="bash"> | ||
| + | cd $HOME/Desktop | ||
| + | mkdir test_dx | ||
| + | cd test_dx | ||
| + | |||
| + | # See sample scripts | ||
| + | ls -la $HOME/software/relax/sample_scripts/model_free | ||
| + | cp $HOME/software/relax/sample_scripts/model_free/map.py . | ||
| + | |||
| + | # Get data | ||
| + | ls -la $HOME/software/relax/test_suite/shared_data/model_free/sphere | ||
| + | cp $HOME/software/relax/test_suite/shared_data/model_free/sphere/*.out . | ||
| + | cp $HOME/software/relax/test_suite/shared_data/model_free/sphere/*.pdb . | ||
| + | |||
| + | # Modify map script | ||
| + | cp map.py map_mod.py | ||
| + | sed -i.bak "s/res_num_col=1)/mol_name_col=1, res_num_col=2, res_name_col=3, spin_num_col=4, spin_name_col=5)/g" map_mod.py | ||
| + | sed -i.bak "s/600/900/g" map_mod.py | ||
| + | sed -i.bak "s/spin\.name/#spin\.name/g" map_mod.py | ||
| + | sed -i.bak "s/res_num_col=1, data_col=3, error_col=4)/mol_name_col=1, res_num_col=2, res_name_col=3, spin_num_col=4, spin_name_col=5, data_col=6, error_col=7)/g" map_mod.py | ||
| + | sed -i.bak "s/sequence\.attach_protons/#sequence\.attach_protons/g" map_mod.py | ||
| + | sed -i.bak "spin/a spin" map_mod.py | ||
| + | |||
| + | # Add these two lines to map_mod.py | ||
| + | spin.element(element='H', spin_id='@H') | ||
| + | spin.isotope('1H', spin_id='@H') | ||
| + | |||
| + | # Run | ||
| + | relax map_mod.py | ||
| + | </source> | ||
| + | |||
== Code to generate map == | == Code to generate map == | ||
<source lang="Python"> | <source lang="Python"> | ||
| Line 8: | Line 42: | ||
for cur_spin, mol_name, resi, resn, spin_id in spin_loop(full_info=True, return_id=True, skip_desel=True): | for cur_spin, mol_name, resi, resn, spin_id in spin_loop(full_info=True, return_id=True, skip_desel=True): | ||
file_name = "map%s" % (cur_spin_id .replace('#', '_').replace(':', '_').replace('@', '_')) | file_name = "map%s" % (cur_spin_id .replace('#', '_').replace(':', '_').replace('@', '_')) | ||
| − | dx.map(params=['dw', 'pA', 'kex'], map_type='Iso3D', spin_id=":1@N", inc=70, lower=None, upper=None, axis_incs=5, file_prefix=file_name, dir=ds.resdir, point=None, point_file='point', remap=None) | + | dx.map(params=['dw', 'pA', 'kex'], map_type='Iso3D', spin_id=":1@N", inc=70, lower=None, upper=None, axis_incs=5, |
| + | file_prefix=file_name, dir=ds.resdir, point=None, point_file='point', remap=None) | ||
#vp_exec: A flag specifying whether to execute the visual program automatically at start-up. | #vp_exec: A flag specifying whether to execute the visual program automatically at start-up. | ||
dx.execute(file_prefix=file_name, dir=ds.resdir, dx_exe='dx', vp_exec=True) | dx.execute(file_prefix=file_name, dir=ds.resdir, dx_exe='dx', vp_exec=True) | ||
| Line 17: | Line 52: | ||
== How to use dx == | == How to use dx == | ||
| − | + | Run <code>dx</code>, and then in the '''Data explorer''' or '''DE'''. | |
| − | + | * Click on <code>Edit Visual Programs...</code>. | |
| − | * Click on | + | * Select the <code>map.net</code> program created by relax, |
| − | * Select the map.net program created by relax, | ||
| − | Now in the | + | Now in the <code>Visual Program Editor</code> or <code>VPE</code>. |
| − | * Select the menu entry | + | * Select the menu entry <code>Execute->Execute on change</code>. |
That's it. | That's it. | ||
| − | You now have a 3D frame, but nothing in it. | + | You now have a 3D frame, but nothing in it. Therefore the contour levels must be too low or high. From the map file, the values are in the hundreds of thousands. |
| − | Therefore the contour levels must be too low or high. | ||
| − | From the map file, the values are in the hundreds of thousands. | ||
Then: | Then: | ||
| − | * In the main program window, double click on the | + | * In the main program window, double click on the <code>Isosurface elements</code>. |
* Change the values until you see surfaces. In the first the value is 500. I changed this to 500,000. Multiply all by 1000. | * Change the values until you see surfaces. In the first the value is 500. I changed this to 500,000. Multiply all by 1000. | ||
* In the second, 100 -> 100000. | * In the second, 100 -> 100000. | ||
| Line 40: | Line 72: | ||
This should maybe be performed by the dx.map user function, determining reasonable contour levels. | This should maybe be performed by the dx.map user function, determining reasonable contour levels. | ||
| − | With a bit of zooming, clicking on | + | With a bit of zooming, clicking on <code>File -> Save image</code> in the <code>Surface</code> window, <code>allowing rendering</code>, and outputting to a large TIFF file, <code>save current</code>, then <code>apply</code>. |
| − | An example image cropped and converted to PNG in the GIMP at | + | An example image cropped and converted to PNG in the GIMP at https://gna.org/bugs/download.php?file_id=20641. |
| − | https://gna.org/bugs/download.php?file_id=20641. | ||
| − | Note that for a good resolution plot, you will need many more increments. | + | Note that for a good resolution plot, you will need many more increments. Using the lower and upper dx.map arguments will be useful to zoom into the space. |
| − | Using the lower and upper dx.map arguments will be useful to zoom into the space. | ||
== Example images == | == Example images == | ||
| − | These images is from | + | These images is from {{gna bug link|22024|text=bug 22024. Minimisation space for CR72 is catastrophic. The chi2 surface over dw and pA is bounded.}} |
{|style="margin: 0 auto;" | {|style="margin: 0 auto;" | ||
| − | | [[File:Bug 22024 R2eff.png|thumb|center|upright=2| | + | | [[File:Bug 22024 R2eff.png|thumb|center|upright=2|{{gna bug url|22024}} Bug 22024: Minimisation space for CR72 is catastrophic. The chi2 surface over dw and pA is bounded.]] |
| − | | [[File:Bug 22024 Dw kex.png|thumb|upright=2| | + | | [[File:Bug 22024 Dw kex.png|thumb|upright=2|{{gna bug url|22024}} Bug 22024: Minimisation space for CR72 is catastrophic. The chi2 surface over dw and pA is bounded.]] |
|} | |} | ||
{|style="margin: 0 auto;" | {|style="margin: 0 auto;" | ||
| − | | [[File:Bug 22024 Dw kex pA.png|thumb|upright=2| | + | | [[File:Bug 22024 Dw kex pA.png|thumb|upright=2|{{gna bug url|22024}} Bug 22024: Minimisation space for CR72 is catastrophic. The chi2 surface over dw and pA is bounded.]] |
| − | | [[File:Bug 22024 Dw pA.png|thumb|upright=2| | + | | [[File:Bug 22024 Dw pA.png|thumb|upright=2|{{gna bug url|22024}} Bug 22024: Minimisation space for CR72 is catastrophic. The chi2 surface over dw and pA is bounded.]] |
| − | | [[File:Bug 22024 Kex pA.png|thumb|upright=2| | + | | [[File:Bug 22024 Kex pA.png|thumb|upright=2|{{gna bug url|22024}} Bug 22024: Minimisation space for CR72 is catastrophic. The chi2 surface over dw and pA is bounded.]] |
|} | |} | ||
| Line 69: | Line 99: | ||
relax cpmg_synthetic.py | relax cpmg_synthetic.py | ||
</source> | </source> | ||
| − | Here is the synthetic data generator code | + | |
| − | === | + | Here is the synthetic data generator code: |
| − | + | ||
| + | {{collapsible script | ||
| + | | type = relax script | ||
| + | | title = Script for calculating synthetics CPMG data. | ||
| + | | lang = python | ||
| + | | script = | ||
############################################################################### | ############################################################################### | ||
# # | # # | ||
| Line 179: | Line 214: | ||
# Add more spins here. | # Add more spins here. | ||
spins = [ | spins = [ | ||
| − | ['Ala', 1, 'N', {'r2': {r20_key_1: 25.0, r20_key_2: 24.0}, 'r2a': {r20_key_1: 25.0, r20_key_2: 24.0}, 'r2b': {r20_key_1: 25.0, r20_key_2: 24.0}, 'kex': 2200.0, 'pA': 0.993, 'dw': | + | ['Ala', 1, 'N', {'r2': {r20_key_1: 25.0, r20_key_2: 24.0}, 'r2a': {r20_key_1: 25.0, r20_key_2: 24.0}, 'r2b': {r20_key_1: 25.0, r20_key_2: 24.0}, 'kex': 2200.0, 'pA': 0.993, 'dw': 0.5} ] |
#['Ala', 2, 'N', {'r2': {r20_key_1: 15.0, r20_key_2: 14.0}, 'r2a': {r20_key_1: 15.0, r20_key_2: 14.0}, 'r2b': {r20_key_1: 15.0, r20_key_2: 14.0}, 'kex': 2200.0, 'pA': 0.993, 'dw': 1.5} ] | #['Ala', 2, 'N', {'r2': {r20_key_1: 15.0, r20_key_2: 14.0}, 'r2a': {r20_key_1: 15.0, r20_key_2: 14.0}, 'r2b': {r20_key_1: 15.0, r20_key_2: 14.0}, 'kex': 2200.0, 'pA': 0.993, 'dw': 1.5} ] | ||
] | ] | ||
| Line 251: | Line 286: | ||
# To map the hypersurface of chi2, when altering kex, dw and pA. | # To map the hypersurface of chi2, when altering kex, dw and pA. | ||
if not hasattr(ds, 'opendx'): | if not hasattr(ds, 'opendx'): | ||
| − | ds.opendx = False | + | #ds.opendx = False |
| − | + | ds.opendx = True | |
# Which parameters to map on chi2 surface. | # Which parameters to map on chi2 surface. | ||
| Line 291: | Line 326: | ||
cur_spin_id = ":%i@%s"%(res_num, spin_name) | cur_spin_id = ":%i@%s"%(res_num, spin_name) | ||
cur_spin = return_spin(cur_spin_id) | cur_spin = return_spin(cur_spin_id) | ||
| + | |||
| + | # Save all the paramers to loop throgh later. | ||
| + | ds.model_analyse_params = cur_spin.params | ||
par_dic = {} | par_dic = {} | ||
| Line 330: | Line 368: | ||
clust_r2 = clust_params[mo_param] | clust_r2 = clust_params[mo_param] | ||
set_r2 = params[mo_param] | set_r2 = params[mo_param] | ||
| − | for key, val in | + | for key, val in min_r2.items(): |
grid_r2_frq = grid_r2[key] | grid_r2_frq = grid_r2[key] | ||
min_r2_frq = min_r2[key] | min_r2_frq = min_r2[key] | ||
| Line 649: | Line 687: | ||
if ds.opendx: | if ds.opendx: | ||
print_res(cur_spins=cur_spins, grid_params=ds.grid_results[i][3], min_params=ds.min_results[i][3], clust_params=ds.clust_results[i][3]) | print_res(cur_spins=cur_spins, grid_params=ds.grid_results[i][3], min_params=ds.min_results[i][3], clust_params=ds.clust_results[i][3]) | ||
| − | + | }} | |
= See also = | = See also = | ||
[[Install dx]] | [[Install dx]] | ||
[[Category:User_functions]] | [[Category:User_functions]] | ||
Latest revision as of 19:44, 16 October 2020
Contents
Sample data to try dx
cd $HOME/Desktop
mkdir test_dx
cd test_dx
# See sample scripts
ls -la $HOME/software/relax/sample_scripts/model_free
cp $HOME/software/relax/sample_scripts/model_free/map.py .
# Get data
ls -la $HOME/software/relax/test_suite/shared_data/model_free/sphere
cp $HOME/software/relax/test_suite/shared_data/model_free/sphere/*.out .
cp $HOME/software/relax/test_suite/shared_data/model_free/sphere/*.pdb .
# Modify map script
cp map.py map_mod.py
sed -i.bak "s/res_num_col=1)/mol_name_col=1, res_num_col=2, res_name_col=3, spin_num_col=4, spin_name_col=5)/g" map_mod.py
sed -i.bak "s/600/900/g" map_mod.py
sed -i.bak "s/spin\.name/#spin\.name/g" map_mod.py
sed -i.bak "s/res_num_col=1, data_col=3, error_col=4)/mol_name_col=1, res_num_col=2, res_name_col=3, spin_num_col=4, spin_name_col=5, data_col=6, error_col=7)/g" map_mod.py
sed -i.bak "s/sequence\.attach_protons/#sequence\.attach_protons/g" map_mod.py
sed -i.bak "spin/a spin" map_mod.py
# Add these two lines to map_mod.py
spin.element(element='H', spin_id='@H')
spin.isotope('1H', spin_id='@H')
# Run
relax map_mod.py
Code to generate map
# See relax help in prompt
help(dx.map)
from pipe_control.mol_res_spin import return_spin, spin_loop
for cur_spin, mol_name, resi, resn, spin_id in spin_loop(full_info=True, return_id=True, skip_desel=True):
file_name = "map%s" % (cur_spin_id .replace('#', '_').replace(':', '_').replace('@', '_'))
dx.map(params=['dw', 'pA', 'kex'], map_type='Iso3D', spin_id=":1@N", inc=70, lower=None, upper=None, axis_incs=5,
file_prefix=file_name, dir=ds.resdir, point=None, point_file='point', remap=None)
#vp_exec: A flag specifying whether to execute the visual program automatically at start-up.
dx.execute(file_prefix=file_name, dir=ds.resdir, dx_exe='dx', vp_exec=True)
How to install dx
See install dx
How to use dx
Run dx, and then in the Data explorer or DE.
- Click on
Edit Visual Programs.... - Select the
map.netprogram created by relax,
Now in the Visual Program Editor or VPE.
- Select the menu entry
Execute->Execute on change.
That's it.
You now have a 3D frame, but nothing in it. Therefore the contour levels must be too low or high. From the map file, the values are in the hundreds of thousands.
Then:
- In the main program window, double click on the
Isosurface elements. - Change the values until you see surfaces. In the first the value is 500. I changed this to 500,000. Multiply all by 1000.
- In the second, 100 -> 100000.
- In the third, 20 -> 20000.
- In the last, 7 -> 7000.
This should maybe be performed by the dx.map user function, determining reasonable contour levels.
With a bit of zooming, clicking on File -> Save image in the Surface window, allowing rendering, and outputting to a large TIFF file, save current, then apply.
An example image cropped and converted to PNG in the GIMP at https://gna.org/bugs/download.php?file_id=20641.
Note that for a good resolution plot, you will need many more increments. Using the lower and upper dx.map arguments will be useful to zoom into the space.
Example images
These images is from bug 22024. Minimisation space for CR72 is catastrophic. The chi2 surface over dw and pA is bounded.
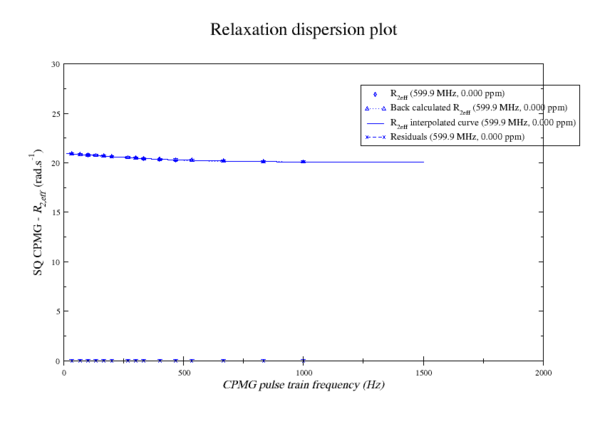 https://web.archive.org/web/gna.org/bugs/?22024 Bug 22024: Minimisation space for CR72 is catastrophic. The chi2 surface over dw and pA is bounded. |
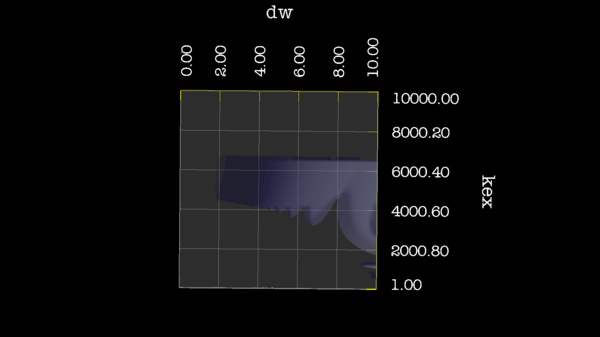 https://web.archive.org/web/gna.org/bugs/?22024 Bug 22024: Minimisation space for CR72 is catastrophic. The chi2 surface over dw and pA is bounded. |
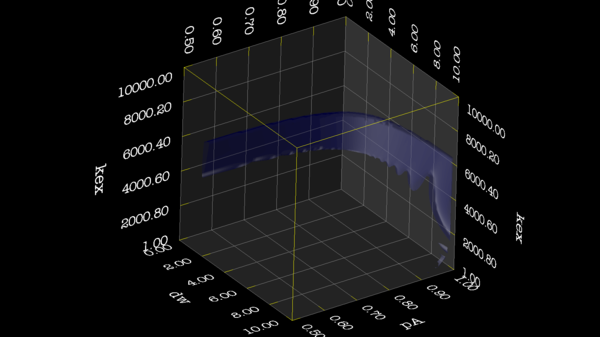 https://web.archive.org/web/gna.org/bugs/?22024 Bug 22024: Minimisation space for CR72 is catastrophic. The chi2 surface over dw and pA is bounded. |
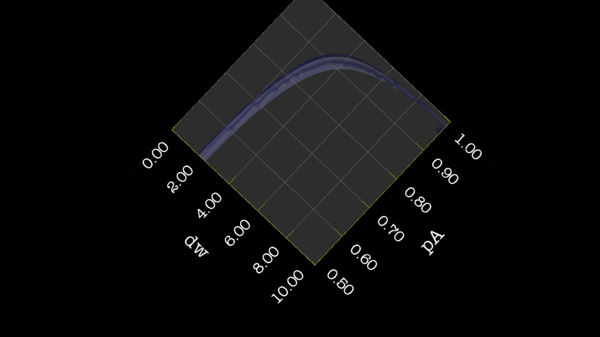 https://web.archive.org/web/gna.org/bugs/?22024 Bug 22024: Minimisation space for CR72 is catastrophic. The chi2 surface over dw and pA is bounded. |
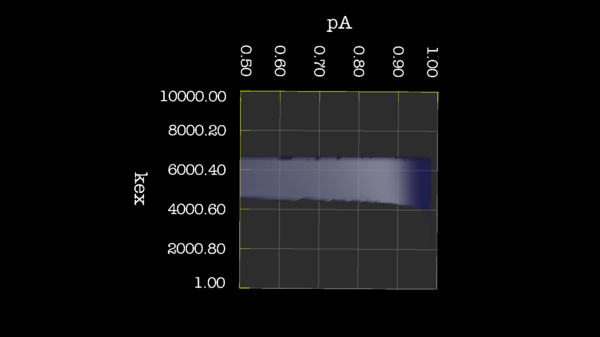 https://web.archive.org/web/gna.org/bugs/?22024 Bug 22024: Minimisation space for CR72 is catastrophic. The chi2 surface over dw and pA is bounded. |
Code to generate map
This script will generate data that can help visualize the different models and the minimisation algorithms in relax.
Call the script cpmg_synthetic.py or similar. Remember .py ending
relax cpmg_synthetic.py
Here is the synthetic data generator code: