Tutorial for sorting data stored as numpy to on-resonance R1rho analysis
Contents
Data background
This is data recorded at 600 and 950 MHz.
This should follow relax-4.0.0.Darwin.dmg installation on a mac, only with GUI.
For each spectrometer frequency, the data is saved in np.arrays
- one for the residue number,
- one for the rates,
- one for the errorbars,
- one for the RF field strength.
They can be retrieved also with scipy's loadmat command.
The experiments are on-resonance R1rho, and the rates are already corrected for the (small) offset effect, using the experimentally determined R1.
Specifically, the numpy shapes of the data is:
- For 600 MHz
- residues (1, 60)
- rates (60, 10)
- errorbars_rate (60, 10)
- RFfields (1, 10)
- For 950 Mhz
- residues (1, 61)
- rates (61, 19)
- errorbars_rate (61, 19)
- RFfields (1, 19)
An example of the data at the 2 fields is:
Create data files for relax
First prepare data, by running in python.
python 1_prepare_data.py
1_prepare_data.py
File: 1_prepare_data.py
import os
import scipy as sc
import scipy.io
import numpy as np
# Set path
cwd = os.getcwd()
fields = [600, 950]
file_names = ['residues', 'rates', 'errorbars_rate', 'RFfields']
# Store data in dictionary
all_data = {}
all_data['fields'] = fields
all_data['file_names'] = file_names
# Make list of residues and make unique
all_res = []
# Loop over the experiments, collect all data
for field in fields:
print "\n", field
# Make a dic inside
all_data['%s'%field] = {}
# Construct the path to the data
path = cwd + os.sep + "Archive" + os.sep + "exp_%s"%field + os.sep + "matrices" + os.sep
all_data['%s'%field]['path'] = path
# Collect all filename paths
field_file_name_paths = []
for file_name in file_names:
# Create path name
file_name_path = path + "%s.mat"%file_name
field_file_name_paths.append(file_name_path)
# Load the data
file_name_path_data = sc.io.loadmat(file_name_path)
# Extract as numpy
file_name_path_data_np = file_name_path_data[file_name]
# And store
all_data['%s'%field]['%s'%file_name] = file_name_path_data
all_data['%s'%field]['np_%s'%file_name] = file_name_path_data_np
print file_name, file_name_path_data_np.shape
# Collect residues
if file_name == "residues":
all_res += list(file_name_path_data_np.flatten())
# Store
all_data['%s'%field]['field_file_name_paths'] = field_file_name_paths
# Make list of residues and make unique
all_res_uniq = sorted(list(set(all_res)))
all_data['all_res_uniq'] = all_res_uniq
# Write a sequence file for relax
f = open("residues.txt", "w")
f.write("# Residue_i\n")
for res in all_res_uniq:
f.write("%s\n"%res)
f.close()
f_exp = open("exp_settings.txt", "w")
f_exp.write("# sfrq_MHz RFfield_kHz file_name\n")
# Then write the files for the rates
for field in all_data['fields']:
resis = all_data['%s'%field]['np_residues'][0]
rates = all_data['%s'%field]['np_rates']
errorbars_rate = all_data['%s'%field]['np_errorbars_rate']
RFfields = all_data['%s'%field]['np_RFfields'][0]
print "\nfield: %3.3f"%field
for i, RF_field_strength_kHz in enumerate(RFfields):
#print "RF_field_strength_kHz: %3.3f"%RF_field_strength_kHz
# Generate file name
f_name = "sfrq_%i_MHz_RFfield_%1.3f_kHz.in"%(field, RF_field_strength_kHz)
cur_file = open(f_name, "w")
cur_file.write("# resi rate rate_err\n")
exp_string = "%11.7f %11.7f %s\n"%(field, RF_field_strength_kHz, f_name)
print exp_string,
f_exp.write(exp_string)
for j, resi in enumerate(resis):
rate = rates[j, i]
error = errorbars_rate[j, i]
string = "%4d %11.7f %11.7f\n"%(resi, rate, error)
cur_file.write(string)
cur_file.close()
f_exp.close()
Run analysis in relax GUI
- Start relax
- Then click: "New analysis (Cmd+n)
- Then click icon for "Relaxation dispersion" -> Next
- Just accept name for the pipe
- Open the interpreter: Cmd+p
- Paste in
import os; os.chdir(os.getenv('HOME') + os.sep + 'Desktop' + os.sep + 'temp'); pwd()
- Then do
script(file='2_load_data.py')
- ( You can scroll through earlier commands with: Cmd+ Arrow up )
- Close the Grace "Results viewer" window
- Close the interpreter window
In the GUI, select:
- For models, only select: "No_Rex" and "M61"
- R1 parameter optimisation to False, ans this on-resonance data does not use R1.
- The number of Monte-Carlo simulations to "3". (The minimum number before relax refuse to run).
- Let other things be standard, and click execute
NOTE:
The number of Monte-Carlo simulations is set to 3.
This means that relax will do the analysis, and the fitted parameters will be correct, but the standard error of the parameters will be wrong.
The reason for this is, that the data "naturally"? for some spins contains measurements which are "doubtful".
When relax is performing Monte-Carlo simulations, all the data is first copied x times to the number of Monte-Carlo simulations, and each R2eff point on the graph
is modified randomly by a gaussian noise with a width described by the associated errors.
For spins which contains measurements which are "doubtful", relax will be spending "very long time" trying to fit a meaningful model.
This "time" is poorly spend on "bad data".
Rather, one should first try to
- analyse the data quickly
- make the graphs
- examine which spins should be deselected
- and then rerun the analysis with a higher number of monte-carlo simulations
After this comes the part with global fit/clustered analysis.
- analyse the data again
- Select which spins should be included in a global fit/clustered analysis
- and then rerun with analysis
2_load_data.py
File: 2_load_data.py
# relax import
from pipe_control.mol_res_spin import spin_loop
# Test if running as script or through GUI.
is_script = False
if not hasattr(cdp, "pipe_type"):
is_script = True
# We need to create a data pipe, which will tell relax which type of data we are expecting
pipe_name = 'relax_disp'
pipe_bundle = 'relax_disp'
pipe.create(pipe_name, pipe_bundle)
# Minimum: Just read the sequence data, but this misses a lot of information.
sequence.read(file='residues.txt', res_num_col=1)
# Name the spins
for cur_spin, mol_name, resi, resn, spin_id in spin_loop(full_info=True, return_id=True, skip_desel=True):
spin.name(name="HN", spin_id=spin_id)
# Manually force the model to be R2eff, so plotting can be performed later
cur_spin.model = "R2eff"
# Name the isotope for field strength scaling.
spin.isotope(isotope='15N')
# Open the settings file
set_file = open("exp_settings.txt")
set_file_lines = set_file.readlines()
for line in set_file_lines:
if "#" in line[0]:
continue
# Get data
field, RF_field_strength_kHz, f_name = line.split()
# Assign data
spec_id = f_name
relax_disp.exp_type(spectrum_id=spec_id, exp_type='R1rho')
# Set the spectrometer frequency
spectrometer.frequency(id=spec_id, frq=float(field), units='MHz')
# Is in kHz, som convert to Hz
#http://wiki.nmr-relax.com/Relax_disp.spin_lock_offset%2Bfield
#http://www.nmr-relax.com/manual/relax_disp_spin_lock_field.html
disp_frq = float(RF_field_strength_kHz)*1000
# Set The spin-lock field strength, nu1, in Hz
relax_disp.spin_lock_field(spectrum_id=spec_id, field=disp_frq)
# Read the R2eff data
relax_disp.r2eff_read(id=spec_id, file=f_name, dir=None, disp_frq=disp_frq, res_num_col=1, data_col=2, error_col=3)
# Is this necessary? The time, in seconds, of the relaxation period.
#relax_disp.relax_time(spectrum_id=spec_id, time=time_sl)
# Plot data
relax_disp.plot_disp_curves(dir='grace', y_axis='r2_eff', x_axis='disp', num_points=1000, extend_hz=500.0, extend_ppm=500.0, interpolate='disp', force=True)
state.save("temp_state", force=True)
# Do it through script
#if False:
#if True:
if is_script:
# Deselect spin 51, due to weid data point
deselect.spin(spin_id=":51@HN", change_all=False)
import os
from auto_analyses import relax_disp as aa_relax_disp
from lib.dispersion.variables import EXP_TYPE_CPMG_DQ, EXP_TYPE_CPMG_MQ, EXP_TYPE_CPMG_PROTON_MQ, EXP_TYPE_CPMG_PROTON_SQ, EXP_TYPE_CPMG_SQ, EXP_TYPE_CPMG_ZQ, EXP_TYPE_LIST, EXP_TYPE_R1RHO, MODEL_B14_FULL, MODEL_CR72, MODEL_CR72_FULL, MODEL_DPL94, MODEL_IT99, MODEL_LIST_ANALYTIC_CPMG, MODEL_LIST_FULL, MODEL_LIST_NUMERIC_CPMG, MODEL_LM63, MODEL_M61, MODEL_M61B, MODEL_MP05, MODEL_NOREX, MODEL_NS_CPMG_2SITE_3D_FULL, MODEL_NS_CPMG_2SITE_EXPANDED, MODEL_NS_CPMG_2SITE_STAR_FULL, MODEL_NS_R1RHO_2SITE, MODEL_NS_R1RHO_3SITE, MODEL_NS_R1RHO_3SITE_LINEAR, MODEL_PARAMS, MODEL_R2EFF, MODEL_TP02, MODEL_TAP03
# Number of grid search increments. If set to None, then the grid search will be turned off and the default parameter values will be used instead.
#GRID_INC = None
GRID_INC = 21
# The number of Monte Carlo simulations to be used for error analysis at the end of the analysis.
MC_NUM = 3
# Model selection technique.
MODSEL = 'AIC'
result_dir_name = os.getcwd()
# Which models to analyse ?
#MODELS = [MODEL_R2EFF, MODEL_NOREX, MODEL_DPL94, MODEL_TP02, MODEL_TAP03, MODEL_MP05, MODEL_NS_R1RHO_2SITE]
#MODELS = [MODEL_R2EFF, MODEL_NOREX]
# R2EFF shall be skipped here
#MODELS = [MODEL_NOREX]
# http://wiki.nmr-relax.com/M61
MODELS = [MODEL_NOREX, MODEL_M61]
# Fit, instead of read
#r1_fit = True
r1_fit = False
# Go
aa_relax_disp.Relax_disp(pipe_name=pipe_name, pipe_bundle=pipe_bundle, results_dir=result_dir_name, models=MODELS, grid_inc=GRID_INC, mc_sim_num=MC_NUM, modsel=MODSEL, r1_fit=r1_fit)
Inspect and make graphs
- When this is done, quit relax
- Then to convert all xmgrace files to png
cd grace
./grace2images.py -t EPS,PNG
This should give the images of the data
- Afterwards you can
- start relax
- Open the interpreter: Cmd+p
- Paste in
import os; os.chdir(os.getenv('HOME') + os.sep + 'Desktop' + os.sep + 'temp'); pwd()
- Close the interpreter
- File -> Open relax state -> "temp_state.bz2"
Here comes an error message we have to investigate. But this can just be closed?
Essentially one need to de-select spin 51 before one select the models for R1rho and then start the analysis.
Run the same analysis in relax through terminal
The analysis can be performed by
python 1_prepare_data.py
relax 2_load_data.py -t log.txt
# Or
mpirun -np 8 relax --multi='mpi4py' 2_load_data.py -t log.txt
Following analysis
First make graphs
cd No_Rex/
./grace2images.py -t EPS,PNG
cat r1rho_prime.out
cd ..
cd M61
./grace2images.py -t PNG
cat kex.out
cat phi_ex.out
cat kex.out | grep "None 12"
cat phi_ex.out | grep "None 12"
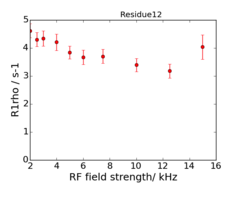
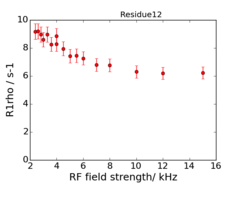
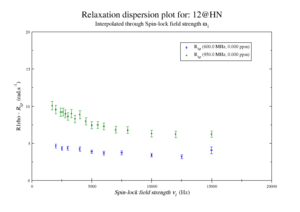
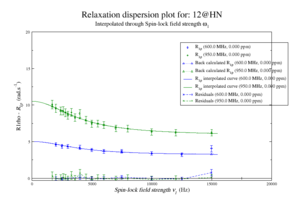
For model M61, spin 12 is:
Resi 12 at 600+950 MHz for M61 model. kex=25000, phi_ex=0.32
Further analysis steps
- Inspect which residues should not be included in analysis
sel_ids = [
":4@HN",
":5@HN",
":6@HN",
":7@HN",
":10@HN",
":11@HN",
]
# De-select for analysis those spins who have bad data
deselect.all()
for sel_spin in sel_ids:
print("Selecting spin %s"%desel_spin)
select.spin(spin_id=desel_spin, change_all=False)
- Inspect which residues should be analysed together for a clustered/global fit.
cluster_ids = [
":13@N",
":15@N",
":16@N",
":25@N"]
# Cluster spins
for curspin in cluster_ids:
print("Adding spin %s to cluster"%curspin)
relax_disp.cluster('model_cluster', curspin)
- Run analysis again