Welcome to the relax Wiki.
| The community-run support site for relax, a program for the study of molecular dynamics using experimental NMR data.
|
User contributions - How to edit pages at the wiki
Please read the guidelines here.
What's new?
|
|
Did you know...
The DPL extension (version 2.3.0) produced a SQL statement which lead to a Database error.
The reason may be an internal error of DPL or an error which you made,
especially when using DPL options like titleregexp.
Query text is:
SELECT DISTINCT `page`.page_namespace AS page_namespace,`page`.page_title AS page_title,`page`.page_id AS page_id, `page`.page_title AS sortkey, `page`.page_counter AS page_counter FROM `page` WHERE 1=1 AND `page`.page_namespace IN ('0') AND `page`.page_is_redirect=0 ORDER BY page_title ASC LIMIT 500 OFFSET 0
Error message is:
Unknown column 'page.page_counter' in 'SELECT' (mysql-n)
|
|
|
Random screenshots
 The analysis selection wizard 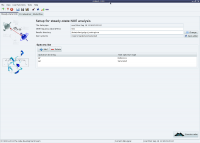 Steady-state NOE analysis
|
|